Markus Schosserer, PhD
Junior PITel.: +43 (0)1 40160 - 56558
E-Mail: markus.schosserer@meduniwien.ac.at
Extension of healthy lifespan
One of the significant challenges modern medicine and biology face today is the continuously increasing life expectance of the population and, consequently, the increase in age-related pathologies. Since the changes in physiology and morphology of organisms, tissues, and cells during their life span are still poorly understood, it is vital to gain insight into the molecular mechanisms of ageing and ageing-associated pathologies to design strategies that allow for maximizing the human health span. Therefore, our research focuses on understanding the cellular and molecular changes that occur during ageing, how these changes affect tissue functionality, and where and how repair and regeneration must counteract these changes (Zeitschrift für Gerontologie und Geriatrie).
Ribosome heterogeneity in ageing
Our lab recently discovered that NSUN5, one of the few highly conserved genes modulating longevity, can influence the ageing process of different organisms. We revealed that protein synthesis in cells could be "reprogrammed" by reducing NSUN5 levels, extending the lifespans of flies, worms and baker's yeast. This study was carried out in cooperation with international partners and published in Nature Communications.
Ribosomes are molecular machines carrying out cellular protein synthesis. In the last few years, it became clear that this is not a purely mechanical process but that differentially composed ribosomes actively regulate it. By building these "specialized" ribosomes, cells are better able to react to different environmental conditions, such as heat, starvation and other types of stress.
NSUN5 adds a single methyl group to ribosomal RNA, one of the ribosome's most essential building blocks. If NSUN5 and thereby the methyl group are missing, these "specialized" ribosomes synthesize proteins, which render flies, worms and yeast more resistant to stress, allowing them to live longer. Thereby we could show for the first time that modification of ribosomal RNA can directly modulate animal lifespan and stress resistance. The perspective of altering the function of a huge molecular machine, like the ribosome, just by a minor modification of a single highly conserved building block (in this case, an RNA nucleotide) and thereby adapting cells to counteract stress better is of high interest, as several mechanisms controlling the life span of organisms converge on the ribosome. Interestingly, we could also demonstrate that loss of NSUN5 in human cells reduces their size, proliferation rate, and body weight (Nucleic Acids Research).
So far, only methylations of DNA were considered to be necessary for the regulation of gene expression. However, this study and others emphasize that methylation of rRNAs, tRNAs, mRNAs and other ncRNAs can impact protein translation and might be involved in post-transcriptional regulation processes. The analysis of RNA methylations at different nucleotides is not yet as well established as the detection of DNA methylations, but new techniques and protocols, such as bisulfite sequencing of RNA for the analysis of m5C methylation, are constantly emerging.
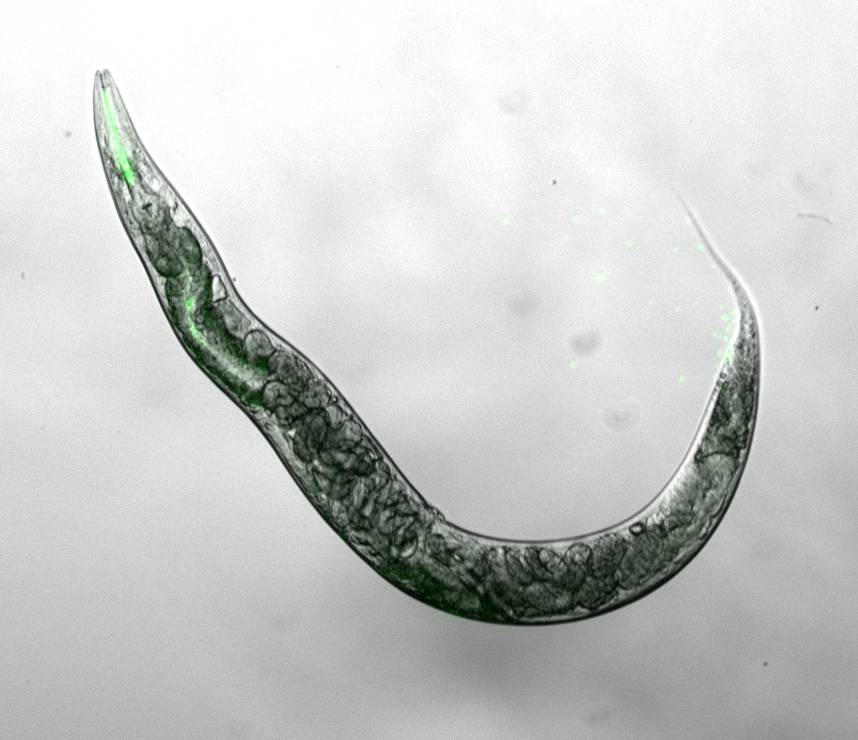
Thus, we aim to identify other ribosomal RNA-modifying enzymes that can also modulate ageing and stress resistance in the nematode Caenorhabditis elegans (Figure 1) and various human cellular ageing models. We have already discovered that NSUN-1 modulates ageing and development in C. elegans (eLife).
Although the gap between simple model organisms and potential applications in humans is significant, we believe that our findings will contribute to a better understanding of evolutionarily conserved ageing processes and related diseases and thereby promote a healthier life in old age. Ageing is one of the significant challenges in our modern society. Therefore, the discovery of novel mechanisms to improve life- and healthspan, like specialized ribosomes (Figure 2), might lead to the development of prognostic markers that help to design intervention steps against the early onset of age-associated diseases.
Cellular senescence and novel models of skin ageing
Together with colleagues at the Department of Dermatology, the Technical University of Vienna and CHANEL Parfums Beauté in the CD-Laboratory "SKINMAGINE", we characterize the effect of plant extracts on cellular senescence and how this might relate to skin functionality in ageing (npj Aging and Mechanisms of Disease).
The human skin is the most exposed organ to factors that accelerate ageing. One factor, cellular senescence, has recently been identified as crucial in tissue's functional decline during ageing (Frontiers in Oncology). It is well established that up to 20% of the skin cells are senescent at an advanced age (Figure 3). However, the influence of senescent cells on the microenvironment in the human skin is not fully understood. Senescent cells acquire a senescence-associated secretory phenotype (SASP) which leads to the secretion of soluble signalling factors and thus changes the micro-environment drastically. Recent studies suggest that microRNAs and lipids are also part of the SASP and can influence the surrounding cells, thus promoting skin ageing (Impact Journals Aging, Journal of Investigative Dermatology, Journal of Investigative Dermatology).

Figure 3: Senescent human dermal fibroblasts stained for senescence-associated beta-Galactosidase activity
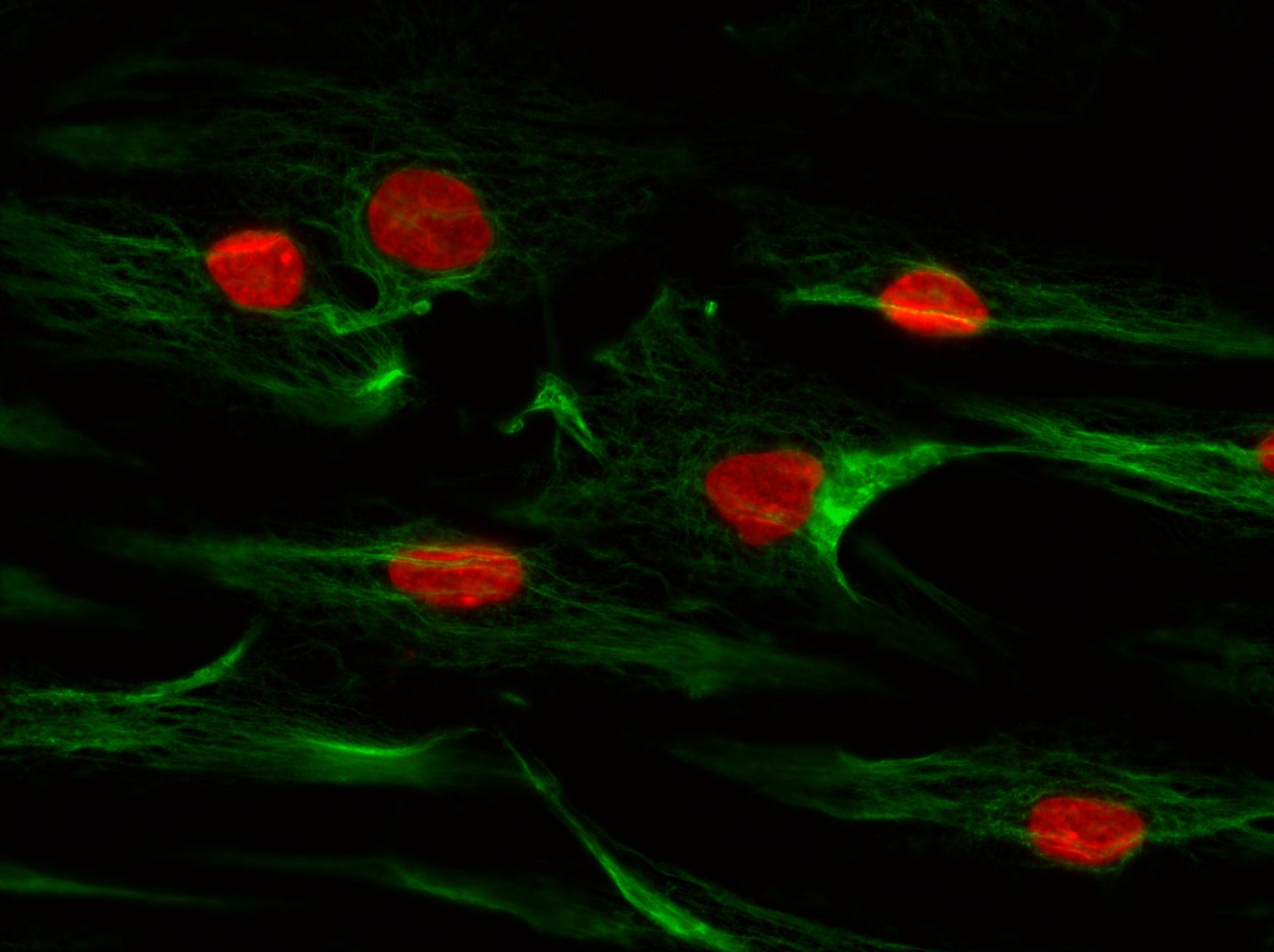
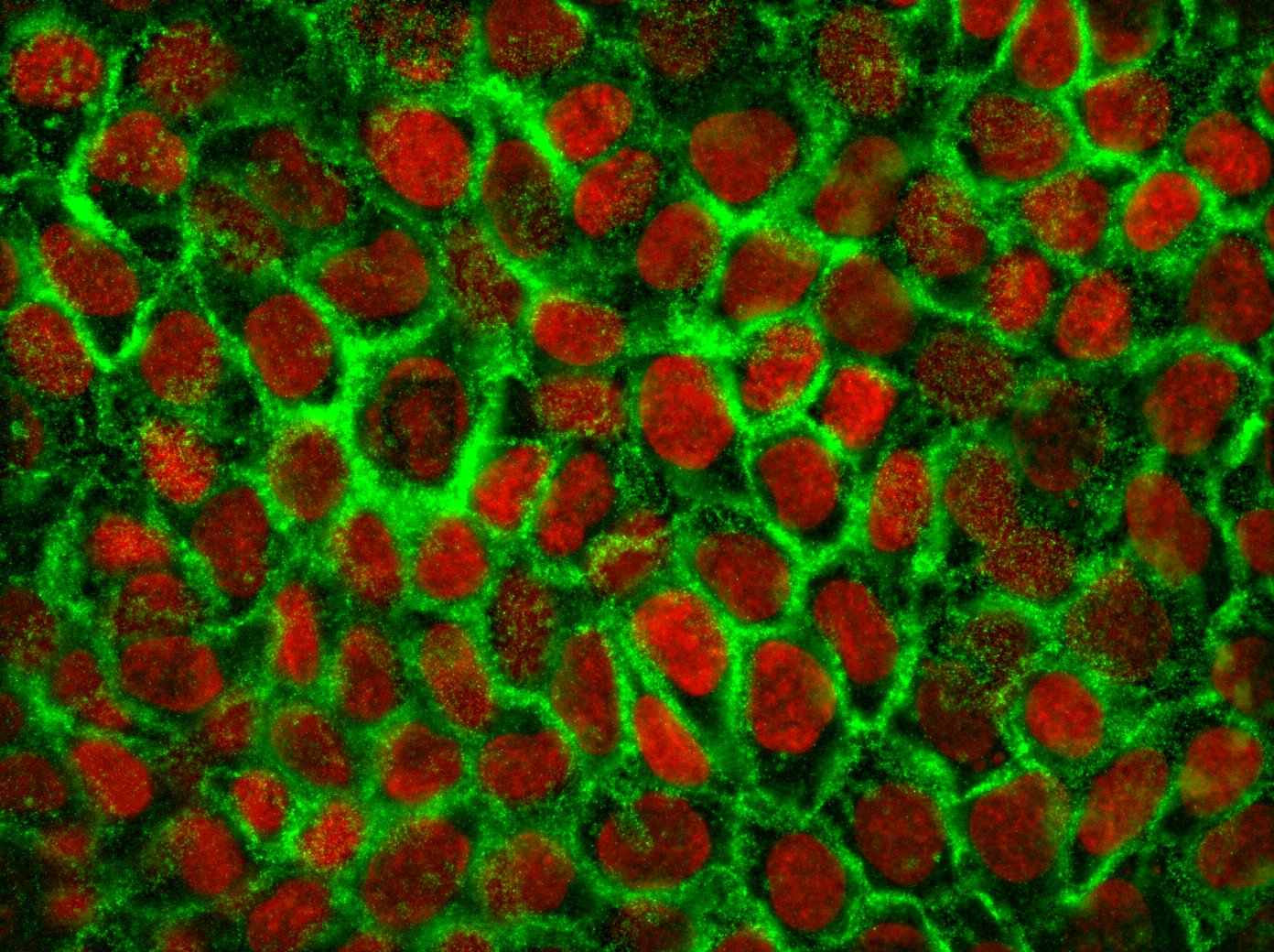
The complex interaction between fibroblasts (Figure 4), extracellular matrix and keratinocytes (Figure 5) cannot be investigated in a two-dimensional cell cultivation system. Thus, developing a robust, three-dimensional "aged" skin equivalent model is required.
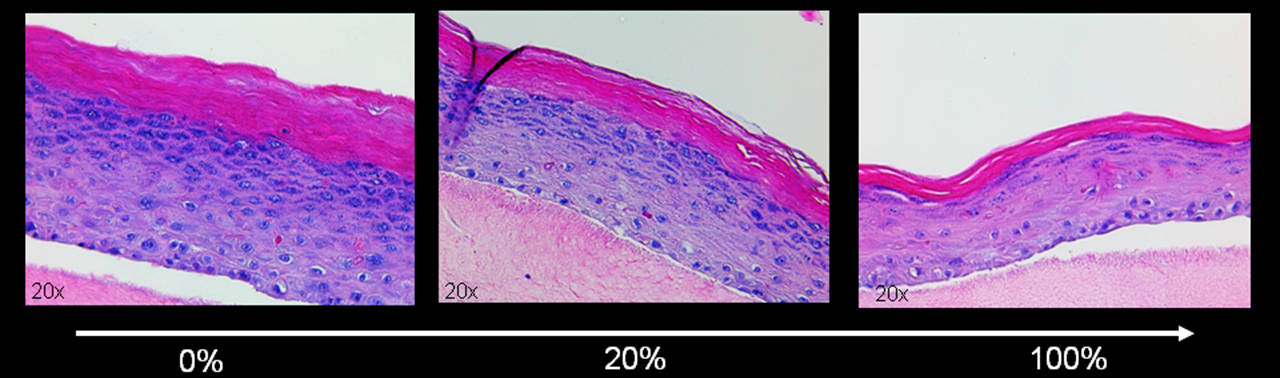
Since fibroblasts in the dermis rarely divide, the cells are more likely to reach senescence by stress-induced events rather than by replication. Hence, we used a stress-induced premature senescent (SIPS) model by repeatedly stressing the cells with H2O2 or doxorubicin. The efficacy of the treatment was verified by two different senescence markers, measurement of growth arrest and staining for senescence-associated beta-galactosidase. The "aged" skin equivalents were then built by adding increasing concentrations of SIPS fibroblasts into the collagen matrix resembling the dermis (Figure 6). Our results suggest that senescent fibroblasts interfere with the differentiation capacity of keratinocytes in the skin equivalent and resemble an "aged" phenotype with thinning of the epidermis and aberrant differentiation of keratinocytes. Additionally, we could show that the senescent cells withstand the stress of being embedded into collagen as good as their non-stressed counterparts and can influence the keratinocytes throughout the whole experiment (npj Aging and Mechanisms of Disease).
These results are promising for further developing a skin equivalent model system with a senescent fibroblast-driven ageing phenotype, thus allowing us to study the crosstalk between senescent fibroblasts and keratinocytes in detail.
Another goal is the development of novel non-invasive, and label-free technologies, such as Raman microspectroscopy, to monitor the causes and consequences of skin ageing.
Markus Schosserer (group leader)
Maximilian Schmid-Siegel (PhD student)
Koray Tav (technician, lab manager)
Matthew Clarke (postdoc)
Anja Wagner (PhD student)
Raul Lopez (PhD student)
Pablo Rubio Bellostas (ERASMUS intern)
Schosserer, M; Minois, N; Angerer, TB; Amring, M; Dellago, H; Harreither, E; Calle-Perez, A; Pircher, A; Gerstl, MP; Pfeifenberger, S; Brandl, C; Mohr, T; Sonntagbauer, M; Kriegner, A; Linder, A; Weinhäusel, A; Steiger, M; Mattanovich, D; Rinnerthaler, M; Karl, T; Sharma, S; Entian, KD; Kos, M; Breitenbach, M; Wilson, IBH; Polacek, N; Grillari-Voglauer, R; Breitenbach-Koller, L; Grillari, J: Methylation of ribosomal RNA by NSUN5 is a conserved mechanism modulating organismal lifespan. Nature Communications 2015; 6/6158; doi: 10.1038/ncomms7158
Heissenberger, C; Rollins, JA; Krammer, TL; Nagelreiter, F; Stocker, I; Wacheul, L; Shpylovyi, A; Tav, K; Snow, S; Grillari, J; Rogers, AN; Lafontaine, DLJ; Schosserer, M: The ribosomal RNA m5C methyltransferase NSUN-1 modulates healthspan and oogenesis in Caenorhabditis elegans. eLife 2020; 9:e56205 doi: 10.7554/eLife.56205
Heissenberger, C; Liendl, L; Nagelreiter, F; Gonskikh, Y; Yang, G; Stelzer, EM; Krammer, TL; Micutkova, L; Vogt, S; Kreil, DP; Sekot, G; Siena, E; Poser, I; Harreither, E; Linder, A; Ehret, V; Helbich, TH; Grillari-Voglauer, R; Jansen-Dürr, P; Koš, M; Polacek, N; Grillari, J; Schosserer, M: Loss of the ribosomal RNA methyltransferase NSUN5 impairs global protein synthesis and normal growth. Nucleic Acids Res. 2019; 47(22):11807-11825, doi: 10.1093/nar/gkz1043
Nagelreiter, F; Coats, MT; Klanert, G; Gludovacz, E; Borth, N; Grillari, J; Schosserer, M: OPP Labeling Enables Total Protein Synthesis Quantification in CHO Production Cell Lines at the Single-Cell Level. Biotechnol J. 2018; Apr;13(4):e1700492; doi: 10.1002/biot.201700492
Wagner, A; Schosserer M: The epitranscriptome in ageing and stress resistance: A systematic review. Ageing Res. Rev. 2022; doi: 10.1016/j.arr.2022.101700
Schosserer, M; Banks, G; Dogan, S; Dungel, P; Fernandes, A; Presen, DM; Matheu, A; Osuchowski, M; Potter, P; Sanfeliu, C; Tuna, BG; Varela-Nieto, I; Bellantuono, I: Modelling physical resilience in ageing mice. Mech Ageing Dev. 2019; 177:91-102; doi: 10.1016/j.mad.2018.10.001
Schosserer, M; Grillari, J; Breitenbach, M: The Dual Role of Cellular Senescence in Developing Tumors and Their Response to Cancer Therapy. Front Oncol. 2017; 7:278; doi: 10.3389/fonc.2017.00278
Liendl, L; Grillari, J; Schosserer, M: Raman fingerprints as promising markers of cellular senescence and aging. Geroscience 2020; 42(2):377-387
Individual working groups:
Ludwig Boltzmann Institute for Experimental and Clinical Traumatology
Ludwig Boltzmann Research Group Senescence and Healing of Wounds
Florian Gruber (Medical University of Vienna, Austria)
Martina Marchetti-Deschmann (Technical University of Vienna, Austria)
Pidder Jansen-Dürr (University of Innsbruck, Austria)
James L. Kirkland / Tamara Tchkonia (Mayo Clinic, USA)
Norbert Polacek (University of Bern, Switzerland)
Denis L.J. Lafontaine (Université Libre de Bruxelles, Belgium)
Aric Rogers / Jarod A. Rollins (MDI Biological Laboratory, USA)
Ilaria Bellantuono (University of Sheffield, UK)
Kostandin Pajcini (University of Illinois at Chicago)
Scientific networks:
Austrian Cluster for Tissue Regeneration
European Epitranscriptomics Network (EPITRAN)
Group of C. elegans new investigators in Europe (GENiE)
European Network for Skin Engineering and Modeling (NETSKINMODELS)
Translational control in Cancer European Network (TRANSLACORE)